Rice University Computer Simulations Explore Molecular Start of Alzheimer’s Disease
Written by |
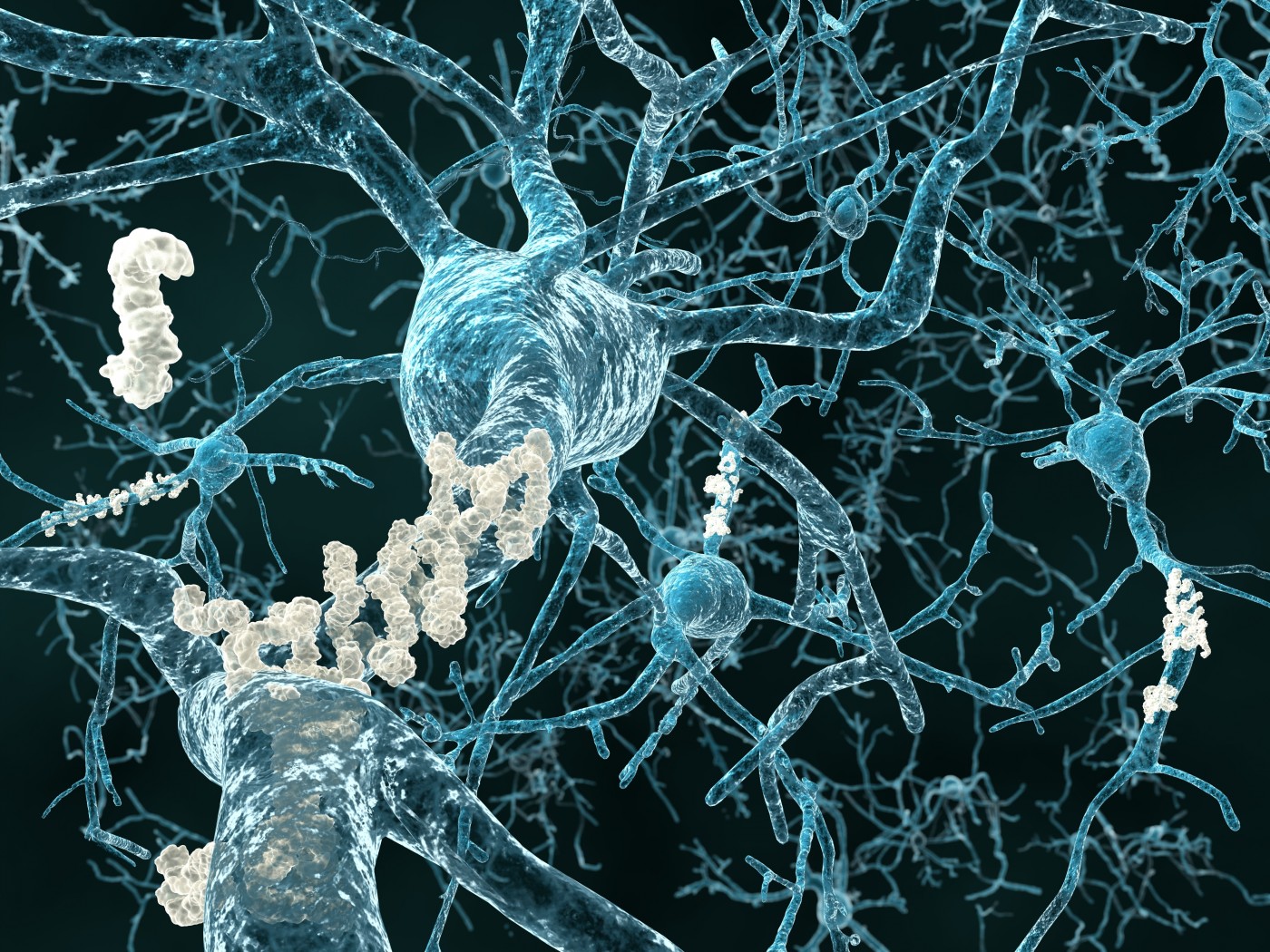
 A new study by researchers at Rice University in Houston used computer simulations to explore the beginnings of the molecular process that marks the start of Alzheimer’s disease, which is triggered by the aggregation of a common protein called amyloid beta. The Rice investigators are the first to model the energy landscape of the pathogenic protein’s assembly of many copies into its toxic form.
A new study by researchers at Rice University in Houston used computer simulations to explore the beginnings of the molecular process that marks the start of Alzheimer’s disease, which is triggered by the aggregation of a common protein called amyloid beta. The Rice investigators are the first to model the energy landscape of the pathogenic protein’s assembly of many copies into its toxic form.

The team of investigators at Rice University who built computer simulations of amyloid beta proteins. From left, Weihua Zheng, Min-Yeh Tsai, Peter Wolynes and Mingchen Chen. Photo credit: Jeff Fitlow/Rice University
The research, led by Prof. Peter Wolynes of Rice’s Center for Theoretical Biological Physics, is reported in the journal Proceedings of the National Academy of Sciences. The study is titled Amyloid Exploring the aggregation free energy landscape of the amyloid-beta protein (1–40).
Wolynes and his team are pioneering the development of the energy landscape theory for proteins, which all start as amino acid chains. The sequence of proteins is determined by DNA, with each arrangement of the chain having a particular associated energy. The mapping of these energies and structures is called the “energy landscape.”
The energy landscape for most proteins guides them to fold into their useful functional shape. “A single molecule of the amyloid beta protein by itself does not fold into a definite structure and also appears to be harmless,” Wolynes said in a press release. “The disease starts when several molecules come together to form an oligomer. The structures of such oligomers, which are thought to be the toxic agents, have been unknown.” At later stage development, the oligomers change into fibrillar plaques that appear to be less toxic.
The Rice researchers used the AWSEM (associative memory, water mediated, structure and energy model), a coarse-grained protein simulation code package being developed and maintained by the the Wolynes lab at Rice and the Papoian lab at the University of Maryland to analyze oligomers of up to eight monomers in the prefibrillar form to see how they change into fibers.
In the oligomer made of six monomers, for instance, they found that this transition takes place when self-recognizing sequences on separate protein molecules interacted with each other rather than with their prefibrillar partners. These structures, they wrote, then form the stable core of a propagating amyloid aggregate.
“A few years ago, there were experiments on the misfolding of a beta protein called titin, and we started to study its aggregation,” Wolynes notes. “We discovered this interesting thing, that it wasn’t the whole protein but only particular parts of the protein that were responsible for misfolding in amyloidogenic regions we found.
“It turned out there were very small lengths of about five amino acids that were extra sticky to each other,” he said. “The surprise was that while the overall molecule was foldable, there were five to 10 residues that, if you had another copy of the same molecule, would be able to stick together and cause aggregation.”
“One problem with previous studies was that the protein concentration changes as the aggregation process proceeds,” Wolynes said, explaining that an important mathematical development using energy landscape theory allowed the Rice researchers to correct for that effect.
 “With that correction, the simulation of oligomers of various complexity predicted solubilities that matched across the board what had been seen in experiments,” said Weihua Zheng, a Rice research scientist, member of the Wolynes lab, and lead author of the study.
“With that correction, the simulation of oligomers of various complexity predicted solubilities that matched across the board what had been seen in experiments,” said Weihua Zheng, a Rice research scientist, member of the Wolynes lab, and lead author of the study.
“One of the reasons you might get Alzheimer’s is that you overproduce this protein,” Zheng said. “That’s why solubility is a key number: You want amyloid beta to remain soluble.”
“The problem may lie in the production of enzymes that keep oligomer concentrations in check,” Wolynes said. “Inhibiting such enzymes is being explored as a route for treatment with drugs.”
“The simulations also demonstrated that the larger the oligomer, the greater the potential for forming fibrils,” Wolynes adding, noting that smaller oligomers of three or four monomers were predicted by the simulations to form cylindrins — which are, as the name suggests, barrel-shaped structures other researchers have suggested could drill into and damage cell membranes. “In future work, we want to look at whether these cylindrical structures actually could penetrate a membrane in simulations,” Wolynes said.
The Rice researchers also noticed that oligomers slightly unfold before going fibrillar just at the point of conversion. “We call that backtracking,” Wolynes said. “It may explain a couple of things, like why these oligomers accumulate rather than all converting to the more benign fibrils.”
The Rice researchers’ simulations, in addition to enabling the mechanism of amyloid beta aggregation to be examined, allowed the lab to analyze the effects of point mutations such as hereditary variants called Dutch and Arctic — that have been associated with early onset Alzheimer’s. In both cases, the mutations seemed to make aggregation faster by exposing hydrophobic segments, though a third mutation, called Iowa, “seems to be mildly protective,” Wolynes said.
In the next stage of their research, the Rice Wolynes team plans to simulate amyloid beta protein aggregation with 42 amino acids for which fiber structures have recently been discovered. “It’s thought that these aggregate more readily than the 40 we studied here, and we hope to see why these are different — or not,” Wolynes said.
Here is a video about the research:
https://www.youtube.com/watch?v=hDWELEEthCs
For more information on Wolynes Research Lab, go to https://wolynes.rice.edu/node/129.
For more information on the Center for Theoretical Biological Physics, go to https://ctbp.rice.edu.